Over a period from 2004 to 2020 I was actively involved full- or part-time in a variety of efforts related to aggregating biomedical and clinical data for use in research networks. These six efforts included:
- Laboratory Animal System (2005-2008)
- NCI Cancer Biomedical Informatics Grid (2004-2010)
- Quantal Universal Exchange Language for Health Information (2010-2012)
- UNC-RTI Self-Generated Health Information Exchange (2012-2015)
- PCORI PCORnet Program (2015-2017)
- Reagan-Udall Foundation for the FDA (2017-2018)
- Consulting on Aggregate Medical Data Networks (2018-2020)
Laboratory Animal System (LASy, 2003-2008)
While working at the Virginia-Maryland Regional College of Veterinary Medicine (VMRCVM) I pursued a research program of my own, gaining an appointment as an adjunct professor in the Department of Biomedical Sciences to pursue these interests. My efforts involved collaboration with two toxicology faculty also in the Department of Biomedical Sciences at VMRCVM, Steve Sheetz in the Department of Accounting and Information Systems in the Pamplin College of Business, and Csaba Egyhazy, in the Department of Computer Science in the College of Arts and Sciences. This developed into various efforts develop to find funding for a prototype Laboratory Animal System (LASy) that used current clinical, toxicology and biomedical information standards (HL7 Relational Information Model, CDISC, from the Clincal Data Interchange Standards Consortium, and NIEHS Toxicology and Biology Data Standards), which could ultimately become a tool for managing and sharing data used by cancer researchers who work with animals. One effort included the creation of a company called BioClinformatics at the VT Corporate Research Center, and obtaining a loan to support the writing of an STTR grant by a professional grant writer. Though this proposal was not funded we did produce a paper about our efforts to produce LASy.1
NCI Cancer Biomedical Informatics Grid – caBIG (2004-2010)
Back in the early days of thinking about how to share biomedical research information in the early 2000’s, one of NCI’s leaders, Ken Buetow wanted to build an integrated set of tools for sharing cancer research information, an initiative that became named the Cancer Biomedical Informatics Grid (caBIG). Around 2004 I saw a presentation by Buetow and decided this was something that I wanted to support, because I considered it the future of biomedical information management. This began my efforts to network with leaders in biomedical informatics, since Buetow as Director of the Center for Biomedical Informatics and Information Technology (CBIIT) had gathered up these people in caBIG as the only major biomedical informatics funding happening – which by 2011 had expended $350 M – which in 2004 was well before this field came into importance with the establishment in 2006 of the NIH Clinical Translational Science Awards (CTSA), providing approximately $500 M in funding to around sixty major academic medical research centers which required integrated biomedical informatics programs to obtain funding.2
Initially I chose to mainly participate in two caBIG workspaces, the Strategic Planning Workspace and the Data Sharing and Intellectual Capital (DSIC) Workspace. Around 2006 I was funded by NCI via Booz Allen Hamilton’s contract as a participant in the DSIC Workspace. The Strategic Planning Workspace, made up of the leaders of all the workspaces, and the NCI caBIG program team, gave me the chance to meet and get to know the people who would become the leaders in biomedical informatics around the country, as well as the program leaders Ken Buetow, Mark Adams, Mary Jo Deering, Chalk Dawson and many others. I got to know several people in the caBIG Strategic Planning Workspace Participants3 including:
Duke Comprehensive Cancer Center, Robert Annechiarico; Fox Chase Cancer Center, Frank J. Manion; Mayo Clinic Comprehensive Cancer Center, Christopher G. Chute; Anderson Cancer Center, Lynn H. Vogel; Ohio State University Research Foundation, Joel H. Saltz; University of Colorado Comprehensive Cancer Center, Jessica Bondy; University of Pennsylvania- Abramson Cancer Centre, David A. Fenstermacher; University of Pittsburgh Medical School, Michael J. Becich.
As the result of my participation in caBIG I also came to know Ulysses Balis, Philip Payne, John Speakman, Howard Bilofsky and many other leaders in biomedical informatics. To build these relationships I participated in monthly workspace calls, several special interest group monthly calls, three face-to-face workspace meetings, as well as the caBIG Annual Meetings between 2005 and 2010. After joining SRI International on their contract with the NIH Center for Information Technology I got involved as project manager for the development of a Clinical Trial Protocol Tracker prototype for the Clinical Trial Management Systems Workspace led by Bob Annechiarico at Duke and John Speakman at NCI. My involvement with caBIG ended in 2010 around the time that caBIG was falling from favor at NCI and shortly after that Ken Buetow left NCI for Arizona State University.
A Quantal Universal Exchange Language – qUEL (2010-2012)
Between full-time positions at SRI International and the School of Information and Library Science (SILS) at the University of North Carolina, Chapel Hill (UNC), I used my connections with leading biomedical informaticists to create the Biomedical Informatics Think Tank (BITT) to support proposals being submitted for the Federal Government’s CIO-SP3, a 10- year Indefinite Delivery/Indefinite Quantity (IDIQ) contract opportunity.
In 2010 a report4 of the President’s Council of Advisors on Science and Technology (PCAST) titled “Realizing the Full Potential of Health Information Technology to Improve Healthcare for Americans: The Path Forward” highlighted major challenges of the current strategy of building a national health information exchange based on funding from the Health Information Technology for Economic and Clinical Health Act (HITECH) that, since 2009, had been motivating a conversion to electronic health records from what had previously been primarily paper-based record system. PCAST recommended the development of a Universal Exchange Language which many of the BITT members wanted to pursue.
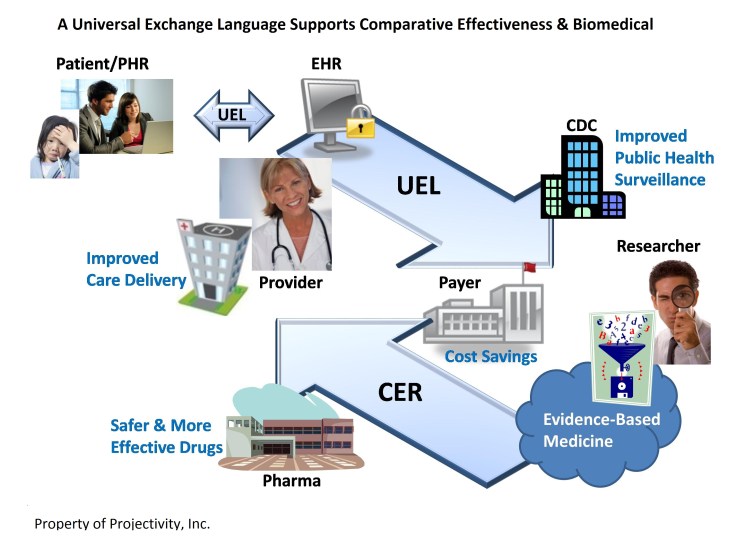
This resulted in the preparation of a white paper5 about the need for a Universal Exchange Language. Then Barry Robson, Ulysses Balis, and myself, developed a prototype Universal Exchange Language which we called Quantal UEL (qUEL) that was built on a unique technology using the methods and tools of quantum mechanics developed by Paul Dirac. Together we produced a method for the secure aggregation model used by quantalUEL6, as well as the specification for quantalUEL7.
In November 2011 Barry Robson and I demonstrated the quantalUEL prototype during the the American Medical Informatics Association (AMIA) Annual Meeting to determine interest in this technology. We also pursued various government sources of funding including SBIR opportunities, and through members of the National Coordination Office (NCO) for the Networking and Information Technology Research and Development (NITRD) Program. We also published an abstract for the International Medical Informatics Association8 and a paper in Computers in Biology and Medicine.9
UNC-RTI Self-Generated Health Information Exchange – SGHIx (2012-2015)
In January 2012, I joined the School of Information and Library Science (SILS) at University of North Carolina (UNC), Chapel Hillto lead a research partnership between SILS and RTI International in health informatics. After 10 months of working to discover an area of mutual interest, leadership of SILS and RTI agreed to pursue the development of research about Self-Generated Health Information. To develop funding for such an effort, I brought together a consortium of companies interested in SGHI, including SGHI Consortium that included representatives from Kaiser Permanente, Alere, Allscripts, Intuit, Blue Cross Blue Shield of NC, Community Care of NC, Johnson & Johnson, Family Health Network, Biz Technology Solutions, Our Health Data Cooperative, Validic, MZI Healthcare, Linguamatics, and IBM. Working with this consortium we defined a need for a Self-Generated Health Information Exchange (SGHIx). I then partnered with Promantus, a consulting company, to develop the web-based SGHIx, test it with users, and present at Health Datapalooza10 and other forums.11,12
PCORI PCORnet Program (2010-2017)
Shortly after the signing of the Affordable Care Act in 2009, which enabled and provided an estimated $2.5 billion for the first ten years of the Patient Centered Outcomes Research Institute (PCORI), I became interested in the possibility that this organization could serve to build an aggregated medical data network to support comparative effectiveness research, since this was the main purpose for pursuing a UEL.5 This led me in 2010 to start attending the monthly calls and face-to-face meetings of the PCORI Board of Governors (BOG). At the May 2011 New York PCORI BOG meeting I made a public comment recommending that PCORI lead in the integration of medical information for increased opportunities for outcomes researchers.13 As expected in July 2012 PCORI led a meeting to discuss the needs for improved electronic medical data14 which led directly to the BOG approving funding for PCORnet, the National Patient-Centered Clinical Research Network. Then between October 2013 and October 2017 I became involved with two parts of the $400 M PCORnet Program:
- A PCORI-Funded PCORnet Patient-Powered Research Network – PPRN (2012-2015)
- The PCORI PCORnet Program (2015-2017).
PCORI-Funded PCORnet Patient-Powered Research Network – PPRN (2012-2015)
In the Fall 2013 a colleague at the School of Medicine UNC Chapel Hill, Michael Kappelman asked me to be the technical project manager on a proposal for a PCORI-funded Crohn’s and Colitis Foundation of America (CCFA) Partners Patient-Powered Research Network (PPRN) he was preparing. Dr. Kappelman thought this effort could benefit from my knowledge and experience working with Validic and the Self-Generated Health Data Exchange (SGHIx). This award was made by PCORI in late 2013 and around March 2014 I was partially funded by this award from PCORI as technical project manager for building the patient portal for the PPRN. We released the PPRN portal for Beta Test in January 2015 and then in full release in March 2015. After about three months we had added approximately another 1000 new participants to the 14,000 previous survey-only study participants. These participants now had an interactive relationship with the researchers giving them the ability to:15
- Create an account and link it to any previous survey information.
- Provide responses to additional surveys.
- Propose, discuss and rank research study suggestions.
- Provide links to their mobile health information (i.e. SGHI), such as from a Fitbit.
- Compare their status on multiple measures with an aggregate of the 15,000 plus participants to determine how well or poorly they were doing in caring for themselves, and how much better they might be doing.
This CCFA Partners PPRN remains in operation as IBD Partners.
PCORI Associate Director, Program Operations, PCORnet Program (2015-2017)
Shortly after the roll-out of the PPRN full release in March 2015, I was offered the position of Director of Program Operations for the $400 M PCORnet program at PCORI in Washington, DC. While at PCORI from June 2015 I helped the PCORnet Program Director reorganize her team and grow it from 8 to 13, while creating an organization that had clear responsibilities, oversight, and relationships with the academic and other private institutions which they were funded through contract awards. Most important, in my judgement, were my efforts to create a master contract – task order mechanism to manage two PCORnet Coordinating Center contracts with Duke/Harvard Pilgrim Health Care, and the Genetic Alliance, contracts that were worth $32 M over a four year period. My master contract approach allowed the division of tasks so that a large contract of this nature could receive detailed management of all milestones.
Reagan-Udall Foundation for the FDA (2017-2018)
I left PCORI in October 2017 to consult full time at Reagan-Udahl Foundation for the FDA which used another medical information-aggregating research network called Sentinel to support research projects of the pharmaceutical industry. I served as project manager for this research network called the Innovation in Medical Evidence Development and Surveillance (IMEDS) Program, managing eight projects with pharmaceutical companies over five months before leaving late February 2018. These companies were using this system to evaluate the safety of medications that were in post-marketing review by the FDA, a critical means for monitoring the actual effects of these treatments in the clinic.
Consulting on Aggregate Medical Data Networks (2018-2020)
Since leaving the Reagan-Udall Foundation I have found opportunities to consult with organizations about connecting to aggregate medical networks including, in addition to PCORnet and IMEDS:
Also I worked with a health care institution on data linkages to the National Death Index and to CMS data.
Footnotes and References
- Sheetz, SD and TP Caruso. Integrating Electronic Health Record Standards into a Laboratory Information Management System. IN Proceedings of the Twelfth Americas Conference on Information Systems, Acapulco, Mexico. August 4-6, 2006.
- “Clinical and Translational Science Award.” In Wikipedia, October 6, 2020.
- caBIG Strategic Planning Workspace. “The Cancer Biomedical Informatics Grid (caBIG): infrastructure and applications for a worldwide research community.” In: Kuhn KA, Warren JR, Leong T-Y, editor. Medinfo 2007: Proceedings of the 12th World Congress on Health (Medical) Informatics: 20–24 August 2007; Brisbane. Amsterdam: IOS Press; 2007. pp. 330–334. [PubMed] [Google Scholar]
- President’s Council of Advisors on Science and Technology (PCAST), White House Office of Science and Technology Policy (OSTP). “Realizing the Full Potential of Health Information Technology to Improve Healthcare for Americans: The Path Forward,” December 8, 2010. Accessed 2020-12-05.
- The Biomedical Informatics Think Tank™. “A Universal Exchange Language Supports Comparative Effectiveness and Biomedical Research.” Feb. 18, 2011. (Available upon request.)
- Robson, B, UGJ Balis, and TP Caruso. “Secure Aggregation Model for a Universal Exchange Language.” The Biomedical Informatics Think Tank™, originally distributed Mar. 25, 2011, updated 10/2011.
- Robson, B. and TP Caruso. “Quantal UEL: A Universal Exchange Language for Health Information – A Specification.” Quantal Semantics, Inc., Aug 13, 2012.
- Robson, B, and TP Caruso. “A Universal Exchange Language for Healthcare.” MedInfo ’13: Proceedings of the 14th World Congress on Medical and Health Informatics, Copenhagen, Denmark, Edited by CU Lehmann, E Ammenwerth, and C Nohr. IOS Press, Washington, DC, USA. Abstract and Poster, August 2013.
- Robson B, Caruso TP, Balis UG. “Suggestions for a web-based universal exchange and inference language for medicine. Continuity of patient care with PCAST disaggregation. Computers in Biology Medicine.” 2015 Jan;56:51-66.
- TP Caruso. A Live App Demo: “Self-Generated Health Information Exchange”, Health Datapalooza 2014. June 2, 2014, Washington, DC.
- TP Caruso. Demo Talk: “Self-Generated Health Information Exchange (SGHIx): Protecting and Providing Benefits for Aggregating Personal Health Data”, 1st International Workshop on Big Data Discovery and Curation at the ASM SIGKDD Knowledge Discovery and Data Mining Conference 2014, New York, NY, August 24, 2014.
- TP Caruso, LK Li, JM Presnell, RG Capra, A Kaul, G Marchionini, and LL Dimitropoulos. “Self‐Generated Health Information Exchange: Providing benefits for sharing personal health data.” Poster Presented AT Health Informatics Career and Internship Fair/Symposium, East Carolina Heart Institute, Greenville, NC, October 24, 2014.
- Barnett, Kerry, Lawrence Becker, Francis Collins, Arnold Epstein, Leah Hole-Curry, Gail Hunt, Robert Jesse, et al. “Board of Governors Meeting Patient-Centered Outcomes Research Institute, May 16, 2011, Public Meeting,” n.d., 5.
- Patient Centered Outcomes Research Institute. “National Workshop to Advance the Use of Electronic Data in Patient-Centered Outcomes Research.” July 2-3, 2012 in Palo Alto, Calif. YouTube Video Link.
- AE Chung, RS Sandler, MD Long, S Ahrens, JL Burris, CF Martin, K Anton, A Robb, TP Caruso, EL Jaeger, W Chen, M Clark, K Myers, A Dobes, MD Kappelman. “Harnessing person-generated health data to accelerate patient-centered outcomes research: the Crohn’s and Colitis Foundation of America PCORnet Patient Powered Research Network (CCFA Partners),” Journal American Medical Information Association, 2016 May;23(3):485-90. Epub 2016 Jan 28.
- Philippakis, Anthony A., Danielle R. Azzariti, Sergi Beltran, Anthony J. Brookes, Catherine A. Brownstein, Michael Brudno, Han G. Brunner, et al. “The Matchmaker Exchange: A Platform for Rare Disease Gene Discovery.” Human Mutation 36, no. 10 (October 2015): 915–21. https://doi.org/10.1002/humu.22858.





